Mechanism of bacterial collagen degradation
Head: Dr. Esther Schönauer
The extracellular matrix (ECM) constitutes one of the first critical physical defence line of the innate immune system against infections in humans and other mammals. Collagens, which are the most abundant and long-lived proteins in humans, form the resilient backbone of the mammalian ECM and ensure the integrity of all our tissues. Collagens are typically resistant to cleavage by most proteases, and collagen remodelling by our endogenous collagenases is tightly regulated and occurs only at specific sites.
For tissue invasion, most pathogens need to cross ECM barriers. Some human pathogens such as Hathewaya histolytica, Clostridium botulinum, Vibrio vulnificus or Leptospira interrogans specifically secrete bacterial collagenases to do so by degrading the collagen scaffold of the ECM. Unlike the endogenous collagenases, these bacterial collagenases can quite radically degrade collagen at multiple sites into small peptides. Thereby, they facilitate the spread of the pathogens into the tissues, promote tissue necrosis and bacterial toxin diffusion and, in the end, aid host infection and colonization.
How bacterial collagenases manage to decompose fibrillar collagen, which is multi-hierarchically organized (fibers, fibrils, tropocollagen, gelatin), is not well understood. Our research was pivotal in identifying the structural enzymatic core required for collagen degradation in bacterial collagenases, i.e. the collagenase unit. The collagenase unit is a two-domain module composed of the activator domain and peptidase domain and it can cleave folded triple-helical collagen in vitro without the aid of other domains.1
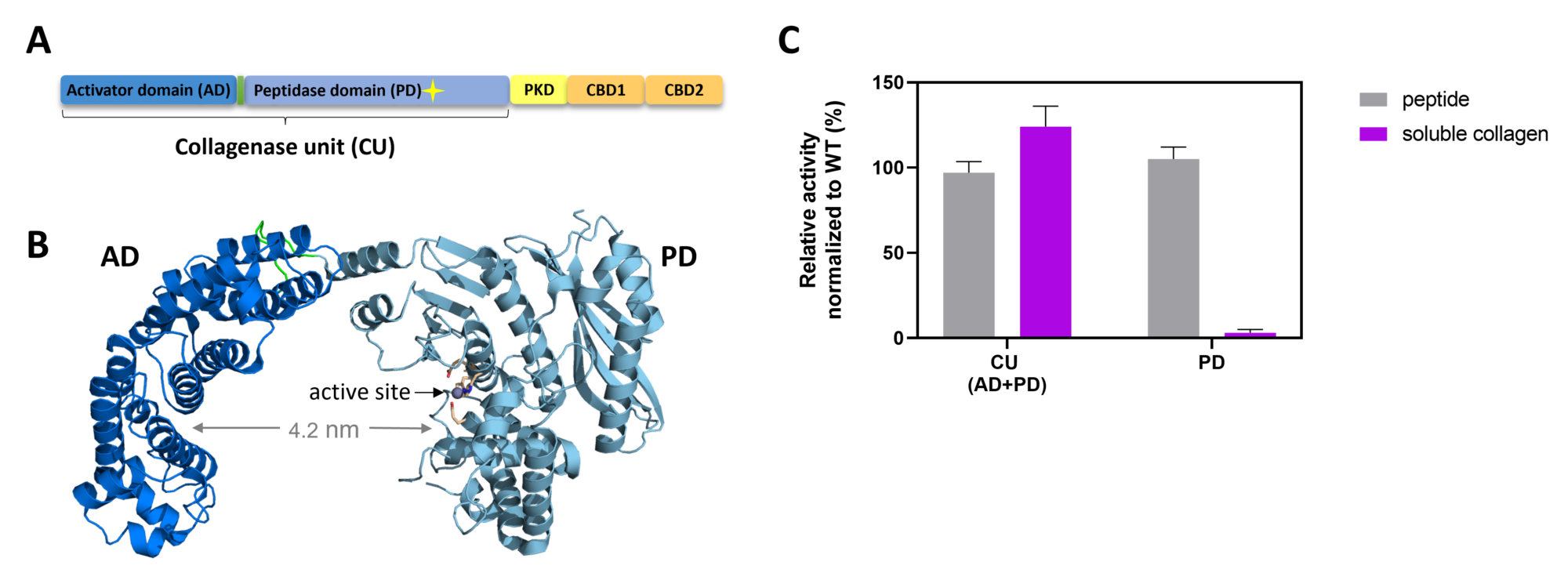
We found that bacterial collagenases have evolved nuanced mechanisms to handle the different forms of collagen (fibrillar vs. soluble), in which, for example, interdomain motions of the AD and PD are instrumental for the processing of fibrillar collagen. Moreover, we revealed that bacterial collagenases use other mechanisms to bind and unfold collagen than human collagenases like MMP-1. In particular, we showed that the presumed non-catalytic activator domain unwinds collagen locally and is required for feeding the unwound substrate to the hydrolytic peptidase domain. Thus, while collagen unwinding is a two-domain job in MMPs, the activator domain in bacterial collagenases can singlehandedly unwind collagen triple helices.2
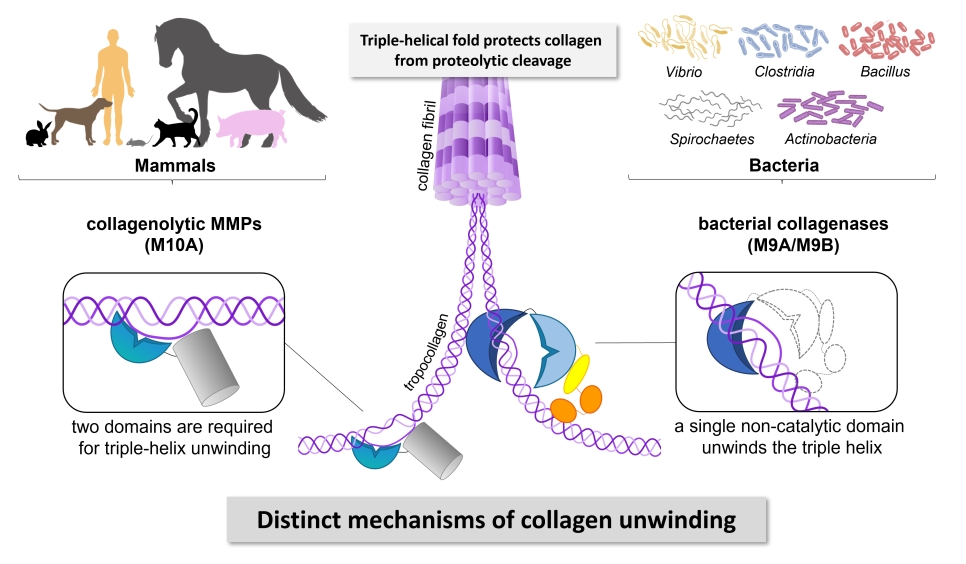
We could trace critical amino acid residues for the binding and unwinding of collagen to a new exosite in the activator domain.2 This discovery paves the way for the development of new selective exosite-directed inhibitors, holding the promise of antivirulence agents for the treatment of infectious diseases with fewer side effects.
Advent of custom-tailored collagenases
Due to their high collagenolytic activity, the bacterial collagenases ColG and ColH from Hathewaya histolytica (formerly Clostridium histolyticum) have found widespread applications in industry, research and clinics. Today, they are routinely used for the isolation of cells from a variety of primary tissues for research and transplantation purposes. In clinics, these bacterial enzymes are used for wound debridement and the treatment of Dupuytren’s contracture and Peyronie’s disease.3–6
Our deepened understanding of the mechanistic details of collagen processing by bacterial collagenases, i.e. deciphering the minimal entities necessary for specific substrate binding, unwinding and cleavage, and the understanding of how domain dynamics affect collagen turnover, have established the foundations to future development of custom-tailored collagenases with unprecedented specificities.
Development of selective active-site directed inhibitors
In collaboration with the Helmholtz Institute for Pharmaceutical Research Saarland (HIPS) (Saarbrücken, Germany) and the Pharmaceutical and Medicinal Chemistry at the Saarland University (Saarbrücken, Germany), we are working on the structure-based design and synthesis of potent and selective inhibitors of clostridial collagenases. Our objective is to generate biologically potent compounds for the future development as anti-infective drug candidates.
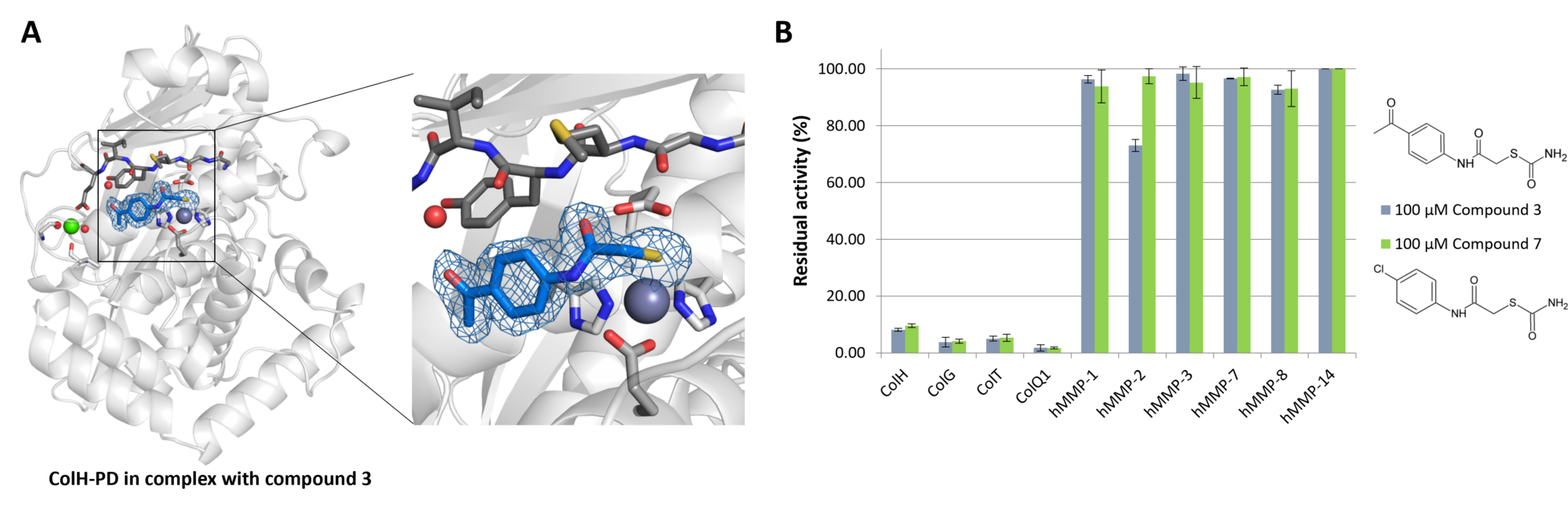

References:
- Eckhard, U., Schönauer, E., Nüss, D. & Brandstetter, H. Structure of collagenase G reveals a chew-and-digest mechanism of bacterial collagenolysis. Nature Structural & Molecular Biology 2011 18:10 18, 1109–1114 (2011). https://www.nature.com/articles/nsmb.2127
- Serwanja, J., Wieland, A. C., Haubenhofer, A., Brandstetter, H. & Schönauer, E. A conserved strategy to attack collagen: The activator domain in bacterial collagenases unwinds triple-helical collagen. Proceedings of the National Academy of Sciences 121, e2321002121 (2024). https://www.pnas.org/doi/10.1073/pnas.2321002121
- Patry, J. & Blanchette, V. Enzymatic debridement with collagenase in wounds and ulcers: a systematic review and meta‐analysis. Int Wound J 14, 1055 (2017).
- Sandler, A. B. et al. Treatment of Dupuytren’s Contracture With Collagenase: A Systematic Review. Hand (N Y) 17, 815–824 (2022).
- Gaffney, C. D. & Kashanian, J. A. Peyronie Disease. JAMA 324, 2566–2566 (2020).
- Ishii, K., Suzuki, N., Mabuchi, Y., Sekiya, I. & Akazawa, C. Technical advantage of recombinant collagenase for isolation of muscle stem cells. Regen Ther 7, 1–7 (2017).
- Schönauer, E. et al. Discovery of a Potent Inhibitor Class with High Selectivity toward Clostridial Collagenases. J Am Chem Soc 139, 12696–12703 (2017). https://pubs.acs.org/doi/full/10.1021/jacs.7b06935





